UW Medicine researchers 'train' artificial intelligence to build vaccines, medications
SEATTLE - Researchers with University of Washington School of Medicine and Harvard University have developed artificial intelligence software to build proteins used in health science.
Their findings were published in the journal Science, explaining that by using machine learning, scientists can "train" artificial intelligence to produce novel solutions to certain prompts. These proteins could potentially be used to make vaccines, cancer treatments or even pull carbon pollution out of the air, their research suggests.
"The proteins we find in nature are amazing molecules, but designed proteins can do so much more," said David Baker, a professor of biochemistry at UW Medicine and senior author on the study. "In this work, we show that machine learning can be used to design proteins with a wide variety of functions."
Scientists ordinarily engineer proteins manually on a computer. These include antibodies and synthetic binding proteins to fight COVID-19, or enzymes for industrial manufacturing.
The problem is that a single protein molecule is already incredibly complex, with thousands of bonded atoms that make them difficult to study, even with specialized software.
That is where AI comes in.
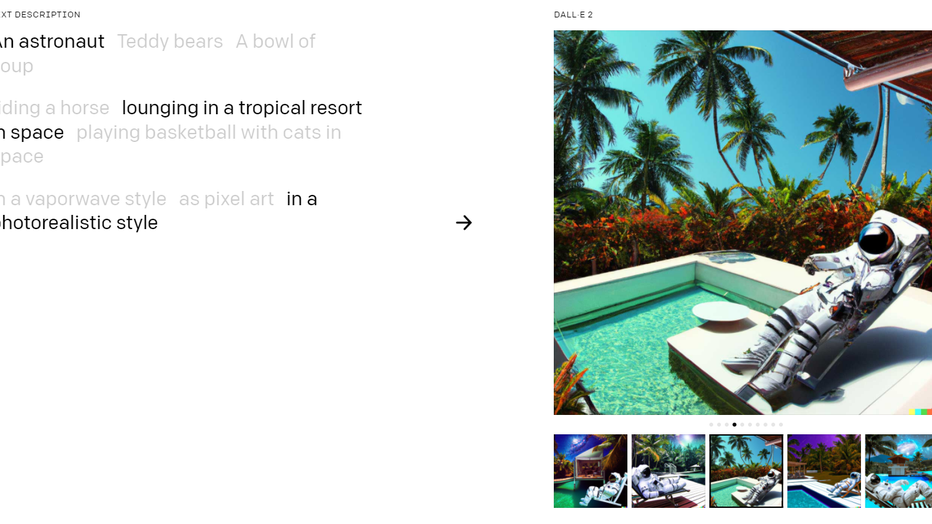
A screenshot from DALL-E 2 demonstrating the AI's ability to generate original images from a prompt (DALL-E 2 // OpenAI)
Several recent projects like DALL-E use text prompts to generate images, which inspired researchers to use the same concept in medicine.
"The idea is the same: neural networks can be trained to see patterns in data," said lead author Joseph Watson, a postdoctoral scholar at UW Medicine. "Once trained, you can give it a prompt and see if it can generate an elegant solution. Often the results are compelling — or even beautiful."
The team "trained" several neural networks with information from the Protein Data Bank—a public database of hundreds of thousands of protein structures from across the animal kingdom.
Researchers say their first approach to generating proteins is dubbed "hallucination," which works like AI image generation tools, which create a new output based on a simple prompt. The second approach is dubbed "inpainting," which works like autocomplete features you might find on search engines or texting.
"Most people can come up with new images of cats or write a paragraph from a prompt if asked, but with protein design, the human brain cannot do what computers now can," said lead author Jue Wang, a postdoctoral scholar at UW Medicine. "Humans just cannot imagine what the solution might look like, but we have set up machines that do."
To generate the protein, the research team compares it to an AI-generated book.
"You start with a random assortment of words — total gibberish. Then you impose a requirement such as that in the opening paragraph, it needs to be a dark and stormy night. Then the computer will change the words one at a time and ask itself ‘Does this make my story make more sense?’ If it does, it keeps the changes until a complete story is written," said Wang.
Rather than an opening paragraph, researchers instead start with a string of amino acids. The software then mutates the sequence over and over until it encodes the desired function, a process that sometimes takes mere seconds.
These sequences can then be manufactured and studied in the lab.
"With autocomplete, or ‘Protein Inpainting,’ we start with the key features we want to see in a new protein, then let the software come up with the rest. Those features can be known binding motifs or even enzyme active sites," said Watson.
According to researchers the proteins made through hallucination and inpainting have been working, including proteins that bind metals, others that bind the anti-cancer receptor PD-1, and still more that could vaccinate against respiratory syncytial virus (RSV).
Extensive testing is needed before these proteins are ever rolled out for medical use.
RELATED: New UW study on effective dispensing of abortion medications via telehealth services
Get breaking news alerts in the FREE FOX 13 Seattle app. Download for Apple iOS or Android. And sign up for BREAKING NEWS emails delivered straight to your inbox.
"These are very powerful new approaches, but there is still much room for improvement," said Baker, who was a recipient of the 2021 Breakthrough Prize in Life Sciences. "Designing high activity enzymes, for example, is still very challenging. But every month our methods just keep getting better! Deep learning transformed protein structure prediction in the past two years, we are now in the midst of a similar transformation of protein design."

